HistonePath ChIP-Seq Service Overview
Active Motif’s HistonePath™ ChIP-Seq Service provides accurate, reproducible identification of histone modification profiles across the genome with high sensitivity and specificity. This technique is widely used in diverse research areas, including stem cell research, studies focused on understanding the mechanisms underlying diseases, and investigating cellular responses to treatments and other stimuli. The HistonePath ChIP-Seq Service can also be useful in identifying new biomarkers since histone modification patterns can be predictive of gene expression and thus be detected prior to changes in gene expression.
Let our team of ChIP-Seq experts help run the experiments and do the bioinformatics for you so you can publish faster and in higher impact journals.
The HistonePath ChIP-Seq Service includes:
- Chromatin preparation and sonication
- Performing ChIP with a ChIP-validated antibody
- Generating ChIP-Seq libraries
- Performing next-generation sequencing
- Comprehensive bioinformatics and data analysis
- Delivery of publication-ready figures
To learn more, please submit an Epigenetic Services Information Request. You can also download Active Motif’s Epigenetic Services Brochure.
HistonePath ChIP-Seq Service Data

Figure 1: ChIP-Seq of small molecular inhibitor-treated cells.
ChIP-Seq was performed from CD34+ cells and antibodies against a histone acetyltransferase (HAT) and its corresponding histone modification. In the presence of a small molecule inhibitor the occupancy of the histone acetyltransferase was specifically reduced at the promoter of the RBBP4 gene but not at the other genes within the 150,000 bp window shown. As expected, the corresponding histone tail acetylation was also reduced at the RBBP4 gene.
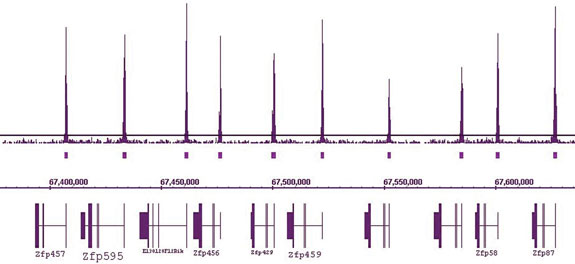
Figure 2: HistonePath™ ChIP-Seq reveals that H3K4me3 peaks are present at the start site of all Zfp genes.
ChIP-Seq was performed using chromatin from mouse livers and a histone H3K4me3 polyclonal antibody (Active Motif cat. no. 39159). The figure focuses on a 3 Mb window containing a Zfp gene cluster on chromosome 13. H3K4me3 peaks are present at the start site of all Zfp genes. Gene annotations run from right to left, therefore the transcription start sites are on the right side of each gene annotation.
HistonePath ChIP-Seq Service Documents
HistonePath ChIP-Seq Service Sample Submission Portal
Our online sample submission portal allows you to easily upload your service project samples and track your project status. Follow the sample submission instructions in the portal to ensure that all your samples arrive at Active Motif in the best possible condition and properly associated with your project.
You might also be interested in:
| Name | Cat No. | Price | |
|---|---|---|---|
| HistonePath™ ChIP-Seq | 25011 | Get Quote | |

