<< Back to MOTIFvations Blog Home Page
Focus on 5hmC: History, Detection, and Applications

By Michelle Tetreault Carlson, Ph.D.
November 9, 2023
Table of Contents:
Introduction
5-hydroxymethylcytosine (5hmC) is a chemically modified form of cytosine, one of several DNA base modifications that can affect gene expression and other cellular processes without changing the fundamental make-up of the four base genetic code. 5hmC is typically discussed in context of 5-methylcytosine (5mC), another cytosine modification which has been studied for many years and is well understood to be associated with repression of transposable elements, X-chromosome inactivation, and genomic imprinting. Loss of this 5mC methylation is controlled both actively and passively by the cell, during the reprogramming that occurs in early embryogenesis, during the formation of memories in neurons, and during the formation of cancers. The first step of active demethylation of 5mC occurs through oxidation by a family of proteins known as the ten-eleven translocation (TET) enzymes to 5hmC. Subsequent oxidation by TET enzymes leads to the formation of 5-formylcytosine (5fC) and 5-carboxylcytosine (5caC), which are then recognized as abnormal and repaired by the cell machinery back to an unmodified cytosine. Originally, it was believed that 5hmC formation was just an intermediate step during this demethylation process. However, studies in the past decade indicate that 5hmC likely not only acts as an intermediate, but also plays an important role in its own right.
Historical Perspective on 5hmC
While the existence of 5hmC has been known since 1952 (Wyatt and Cohen) not a lot of attention was paid to this modification until 2009 when two publications inspired renewed interest. One showed its abundance (0.2-0.6%) in the nuclear DNA of human and mouse brains (Kriaucionis & Heintz). The other showed its abundance in embryonic stem cells and identified TET1 (Ten Eleven Translocation 1) as the enzyme which converts 5mC to 5hmC (Tahiliani et al.). Then, in 2014, it was demonstrated that most 5hmC modifications in the brain are in fact stable throughout the cell life cycle, giving greater support to its role as a true epigenetic mark (Bachman et al.). Much has been learned recently about the distribution and context of 5hmC in the genome. In mammalian DNA, 5hmC is predominately found in CpGs, with only a very small percentage (~2.5%) found in non-CpG context (Schutsky et al.). The distribution pattern of 5hmC shows highest levels in the gene bodies of transcriptionally active genes (He et al.). In contrast to 5mC, the abundance of 5hmC varies significantly between tissues, from highs of 0.5-0.6% in brain, rectum, and colon to lows of 0.05%-0.5% in heart, breast, and placenta (Li et al.). Moreover, around one-third of 5hmC peaks are tissue-specific, and potentially regulate the expression of nearby tissue-specific functional genes (He et al.).
Understanding the molecular mechanisms controlling the role of 5hmC in gene regulation is still in early stages, but it’s clear that aberrant 5hmC patterns are linked to numerous diseases. Dysregulation of 5hmC has been associated with various neurological disorders including Rett syndrome (Brown et al.), autism (Cheng et al.), and depression (Gross et al.). It’s also well known to be involved in cancer, where global loss of 5hmC is a hallmark (Jeschke et al.).
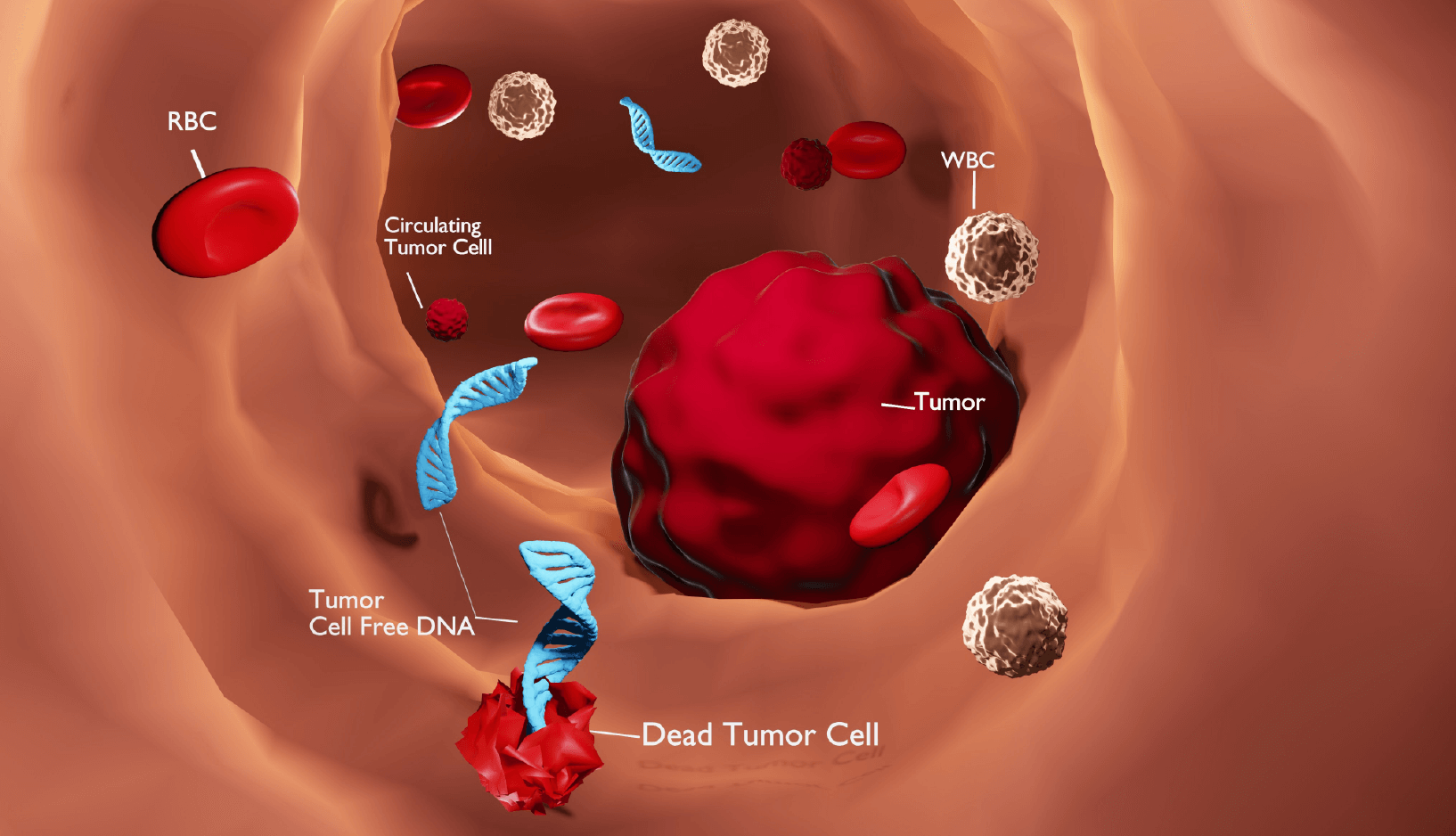
It’s the presence or absence of these specific 5hmC modifications, along with its high tissue specificity that gives 5hmC the potential to serve as an excellent biomarker for disease. Patterns of 5hmC methylation in blood cell-free DNA are especially being explored as potential non-invasive diagnostic and prognostic biomarkers.
Mapping 5hmC
There are numerous methods for identifying and quantifying levels of 5hmC in DNA from tissues and cells, each one with its own advantages and limitations.
Standard chemical-based techniques for determining methylation status at single base resolution, like whole genome bisulfite sequencing (WGBS), cannot distinguish 5hmC from 5mC, as they both respond the same way to bisulfite treatment. The same is true when using standard enzymatic based techniques such as TET-assisted pyridine borane sequencing (TAPS) and enzymatic methyl-seq (EM-seq) (Liu et al., Vasisvil et al.).
To differentiate 5hmC from 5mC in DNA, some clever tricks need to be employed. In oxidative bisulfite sequencing (oxBS-seq) an additional step first oxidizes 5hmC to 5-formylcytosine (5fC) before bisulfite treatment making it sensitive to deamination by bisulfite. With this extra step, the position of all 5mCs can be determined. To then deduce the position of 5hmC, a second standard bisulfite conversion is run in parallel and compared to the first. One of the drawbacks of OxBS-seq is the sequencing depth required, along with a compounding of errors due to having to do two sequencing runs (Booth et al.).
Two more direct methods for determining 5hmC are Tet-assisted bisulfite sequencing (TAB-seq) and a variation of TET-assisted pyridine borane sequencing (TAPS) known as TAPSβ. With TAB-seq 5hmC is first altered by glycosylation. The DNA is then treated with a TET enzyme that converts the 5mC to 5caC but not the glycosylated 5hmC. Subsequent bisulfite treatment only deaminates the 5caC and unmethylated cytosines while the glycosylated 5hmC remains unaffected (Yu et al.). In a similar fashion, for TAPSβ the 5hmC is glycosylated before treatment with a TET enzyme, protecting it from conversion with pyridine borane (Liu et al.). While both TAB-seq and TAPSβ are more direct methods than OxBS-seq for determining which cytosines are hydroxymethylated, they also both rely on the activity of TET enzymes which may not always be 100% efficient and can be expensive to make.
Another bisulfite-free method for localizing 5hmC at single base resolution is a technique developed in Hao Wu’s and Rahul Kohi’s lab named APOBEC-coupled epigenetic sequencing (ACE-seq). ACE-seq uses the APOBEC3A (A3A) enzyme, which can catalyze the deamination of both unmethylated and methylated cytosines to uracil, while leaving 5hmC unconverted. Sequencing can then be used to directly identify 5hmC (Wang et al.). ACE-seq shows results that are in good agreement with other methods such as TAB-seq and OxBS-seq but requires less input.
While determining the genome wide location of 5hmC at single base resolution gives the most information, methods for doing so are also currently the most expensive, time consuming, and require the most sample input. Therefore, enrichment-based methods, which first selectively narrow down the sample to methylated regions, are more widely applied for large scale studies.
One such assay, Hydroxymethylation DNA Immunoprecipitation (hMeDIP), is an immunocapture technique where an antibody specific for hydroxymethylated cytosines is used to immunoprecipitate DNA fragments containing hydroxymethylated genomic DNA (Nester et al.). The enriched DNA can then be used to look at hydroxymethylation inside a particular locus by qPCR or for preparing DNA libraries for gene-wide sequencing. The downside of immunoprecipitation is it’s only as good as the quality of the antibody and the stringency of the binding conditions. While high stringency conditions can eliminate weak binding of 5-hmC, low stringency conditions can increase non-specific binding, giving a higher background. An advantage of the antibody method is that all hydroxymethylated cytosines can be targeted regardless of whether they are in a CpG context. However, this is a very small percentage of hydroxymethylation in mammals.
Another technique, referred to as 5hmC-Seal, also enriches hydroxymethylated DNA fragments. But instead of an antibody, it uses the β-glucosyltransferase (β-GT) enzyme. The β-GT enzyme selectively modifies 5hmC using a UDP-glucose donor to attach a glucose moiety to 5hmC (Song et al.). This is followed by biotinylation of the sugar and subsequent capture of 5hmC containing DNA with streptavidin magnetic beads. The strong binding properties of biotin-streptavidin enable high stringency washes without sacrificing the sensitivity of the assay. This makes the assay less sensitive to stringency issues than an antibody-based method.
Using 5hmC as a Biomarker
During the past several decades, there has been a growing interest in developing methods for both early detection of and monitoring treatment response to cancer. This is especially true for those cancers which are typically not evident until advanced stages such as pancreatic, ovarian, or liver. Early focus in this area has been on detecting proteins, altered gene expression and genetic mutations. More recently, attention is being turned to looking for altered DNA methylation patterns in blood as a screening method, as carcinogenesis is typically accompanied by widespread DNA methylation changes in the cell. Methylation based biomarkers may offer advantages over RNA or genetic biomarkers as they are more stable in bodily fluids (Oliver et al.). And while the abundance of 5mC can be 14 times higher than for 5hmC in human DNA, 5hmC has higher tissue-specificity, which may make it easier to determine cancer origin. Loss of 5hmC methylation in gene promoters and gene bodies has been observed in numerous malignancies, with levels of 5hmC reduced 50-90% in lung cancer, colorectal cancer, glioblastoma, gastric cancer, liver cancer, and malignant melanoma (Bisht et al.).
The best method for mapping and monitoring 5hmC as a biomarker is going to be one that’s sensitive, robust, and requires low DNA input. The 5hmC-Seal technology is proving to be one such method, with numerous publications using this technique to look for biomarkers in circulating cell free DNA (Xu et al.). 5hmC-Seal was used for genome-wide profiling of both cfDNA and in gDNA of paired tumor and adjacent tissues collected from 260 patients recently diagnosed with colorectal, gastric, pancreatic, liver, or thyroid cancer and compared to that from 90 healthy individuals. The results showed robust cancer associated 5hmC patterns in cfDNA which were characteristic of specific cancer types. 5hmC based biomarkers were especially predictive of colorectal and gastric cancers (Li et al.).
In another study 5hmC was mapped in using 5hmC-Seal in cfDNA from 73 newly diagnosed patients with different subtypes of non-Hodgkin lymphoma. Nearly 300 differently modified genes were determined between diffuse large B-cell lymphoma (DLBCL) and follicular lymphoma (FL). 5hmC differences in just four of these genes could be used to distinguish the two cancers in 89% of the patients (Chiu et al.).
The same technique was used in a study to profile differences in 5hmC between patients with acute myeloid leukemia (AML) and non-cancer controls. This allowed the authors to develop a diagnostic model that differentiated AML patients from healthy controls with high sensitivity and specificity. They also developed a prognostic model that predicted prognosis in AML patients (Shao et al.).
In 2020, Chuan He’s lab used 5hmC-Seal to create a human tissue map of 5hmC in 19 human tissues derived from ten organ systems. This was created with the idea that it could serve as a resource to facilitate future studies for understanding the pathogenesis of 5hmC in disease and its development for biomarkers (Xiao-Long Cui et al.).
Finally, one company, ClearNote Health (formally Bluestar Genomics) has successfully used the 5hmC-Seal method to create a noninvasive pancreatic cancer test using cfDNA. In 2021 the test received FDA Breakthrough Device designation for its test in patients with new-onset diabetes. Looking at epigenomics and genomic profiles, with just a simple blood test it detects pancreatic cancer with a sensitivity of 67% and a specificity of 97% in these high-risk patients.
Conclusion
The discovery and understanding of 5-hydroxymethylcytosine (5hmC) as an epigenetic modification have evolved over several decades. A series of key studies published in 2009 and 2010 demonstrated that 5hmC is abundant in certain tissues, particularly in the brain, and is involved in gene regulation and development. It has become increasingly clear that 5hmC is not just an intermediate in DNA demethylation but also has its own distinct epigenetic regulatory functions. In the 2010s, researchers developed and refined techniques such oxBS-seq, TAB-seq, ACE-seq and 5hmC -Seal for mapping 5hmC on a genome-wide scale, enabling comprehensive profiling of 5hmC in various cell types and tissues.
With these advances, our understanding of the significance of 5hmC as an epigenetic mark continues to grow, potentially opening new avenues for therapeutic interventions and disease management. 5hmC-seal, due to its robustness, low input requirements, and genome wide mapping of 5hmC patterns, is one tool successfully being used to determine biomarkers in cell-free DNA, with several studies already showing it to be an effective method to detect and distinguish different types of cancer.
About the author

Michelle Tetreault Carlson, Ph.D.
Michelle’s interest in science was first spurred by the starry skies above her rural farm in upstate New York State, leading her to pursue a B.S. in physics. She was originally interested in astrophysics when entering the University of California, San Diego, but transitioned towards the more practical pursuit of biology earning her Ph.D. in Biophysics, studying photosynthetic proteins. Michelle’s postdoctoral research on retinal ion channels, took her further towards biology, ultimately leading to a career in the biotech industry. She enjoys chatting with scientists about their projects and interacts with them both as a Technical Support Scientist and Product Manager for Active Motif’s DNA Methylation products.
Michelle is a mother of 4 kids and 2 cats, and her hobbies include puzzles (the sign of a patient and logical mind), cooking, and pondering the human condition.
Contact Michelle with any questions at [email protected]
Related Articles
DNA Methylation’s Impact on cfDNA Fragmentation – Considerations When Choosing Biomarkers
August 29, 2023
A new understanding of how differential DNA methylation may impact cell-free DNA fragmentation may help in the identification of powerful biomarkers in liquid biopsies .
Read More
DNA Methylation and Aging: Best of Friends, Worst of Enemies?
January 10, 2023
New research into links between DNA methylation, aging clocks, and Alzheimer’s disease. We look at two recent studies involving epigenetic clocks and epigenetic editing via CRISPR/Cas9 that further demonstrate the link between DNA methylation alterations and normal human aging.
Read More
DNA Methylation for Predicting Prostate Cancer: Where Do We Stand?
December 13, 2022
Some say that Epigenetics touches everything. The marriage between DNA methylation, one of the earliest and most studied of epigenetic mechanisms, and its potential to be a powerful biomarker for disease is exciting and promising. In this overview, we look at prostate cancer research in particular - where we stand in this regard, and also a “new” biomarker associated with voltage-dependent calcium channels!
Read More
<< Back to MOTIFvations Blog Home Page