<< Back to MOTIFvations Blog Home Page
Comprehensive Guide to Understanding and Using CUT&Tag Assays
By Anne-Sophie Ay-Berthomieu, Ph.D.
November 24, 2020
The chromatin immunoprecipitation (ChIP) assay is the gold standard technique to analyze the binding of transcription factors and cofactors to DNA and the localization of histones and histone modifications throughout the genome.
Chromatin immunoprecipitation protocols involve a multi-step process where each step is of crucial importance to obtain interpretable and reproducible ChIP results that can be trusted. Furthermore, ChIP assays generally require a relatively high amount of starting material, which is difficult to obtain for some sample types. Although the read-out for ChIP-Seq assays has become more efficient as a result of improvements to next-generation sequencing technology in recent years, the core principles of the chromatin immunoprecipitation protocol itself have not significantly evolved since it was first developed over 30 years ago.
While the ChIP assay is still the primary method to investigate protein-chromatin interactions, alternative strategies are being developed to overcome some of the limitations of ChIP assays.
In particular, more and more scientists are starting to use Cleavage Under Targets and Tagmentation (CUT&Tag) technology to investigate genomic localization of some histone modifications and transcription factors, especially with low cell numbers.
The CUT&Tag method is a variation of Active Motif’s patented TAM-ChIP™ technology. CUT&Tag is based on the main ChIP principles, antibody-based binding of the target protein or histone modification of interest, but instead of an immunoprecipitation step, antibody incubation is directly followed by the shearing of the chromatin and library preparation. CUT&Tag assays take advantage of a Tn5 transposase that is fused with protein A to direct the enzyme to the antibody bound to its target on chromatin. The Tn5 transposase is pre-loaded with sequencing adapters (generating the assembled pA-Tn5 adapter transposome) to carry out antibody-targeted tagmentation.
CUT&Tag shows a lot of promise and has the potential to alleviate some ChIP limitations, but it also has its own set of limitations that must be considered. This article covers what CUT&Tag is and describes the advantages and drawbacks of this method.
What is CUT&Tag?
CUT&Tag, which is short for Cleavage Under Targets and Tagmentation, is a molecular biology method that researchers use to investigate interactions between proteins and DNA and to identify DNA binding sites for their protein of interest.
Although CUT&Tag is similar in some ways to ChIP assays, the starting material for CUT&Tag is live permeabilized cells or isolated nuclei rather than the cells or tissue that are crosslinked with formaldehyde that are used in ChIP.
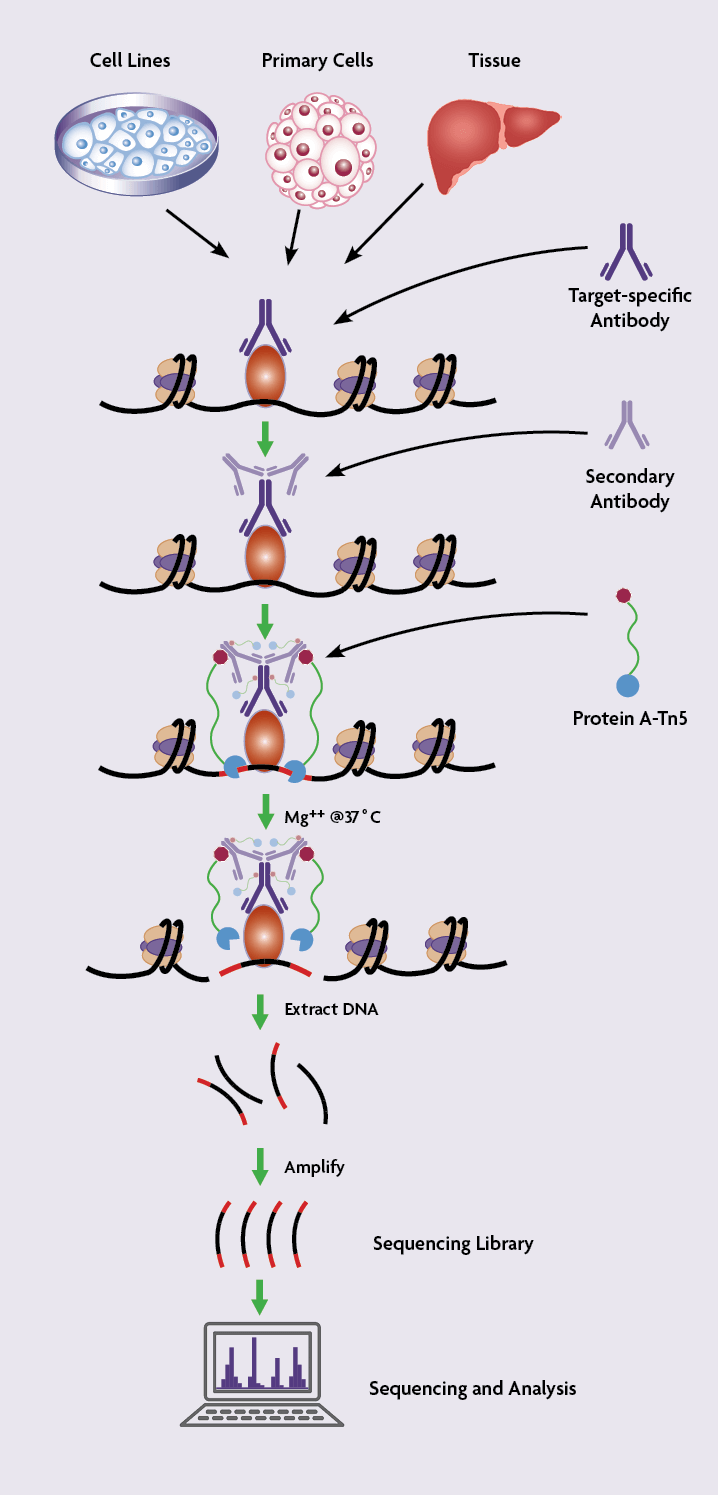
In CUT&Tag protocols, cells are first permeabilized and incubated with an antibody immobilized on concanavalin A-coated magnetic beads to facilitate the subsequent washing steps. Next, the cells are incubated with a primary antibody specific for the target protein of interest followed by incubation with a secondary antibody. The cells are then incubated with assembled transposomes, which consist of protein A fused to the Tn5 transposase enzyme that is conjugated to NGS adapters. After the incubation, unbound transposome is washed away using stringent conditions.
Tn5 is an Mg2+-dependent enzyme so Mg2+ is added to activate the reaction, which results in the chromatin being cut close to the protein binding site and simultaneous addition of the NGS adapter DNA sequences. This leads to chromatin cleavage and library preparation in one single step.
The CUT&Tag method is very sensitive, it has been reported to work with as few as 60 cells for some histone modifications.
With ChIP, sonication randomly shears chromatin and the antibody immunoprecipitates DNA with different lengths, usually several hundred base pairs in length. However, with CUT&Tag, the transposase only cuts chromatin at close proximity to the protein binding site, resulting in shorter lengths of DNA being sequenced. This allows lower sequencing depth (3-5 million reads) to generate robust data, with lower background signal than most ChIP-Seq assays.
Finally, because the CUT&Tag protocol uses intact cells as the starting material, rather than sonicated chromatin, it can be adapted to single-cell experiments (scCUT&Tag).
A Brief History of CUT&Tag and Similar Methods
Many different approaches have been developed to try to improve on crosslinking ChIP (X-ChIP) protocols to produce higher quality results from lower amounts of starting material. In particular, some methods such as Chromatin Endogenous Cleavage (ChEC) and DamID are based on the tethering of nuclease or DNA modifying enzymes to a DNA-binding protein, where the enzyme digests or modifies DNA in the local vicinity of the binding site.
DamID and ChEC methods are based on taking advantage of the DNA adenine methyltransferase enzyme and the micrococcal nuclease (MNase) enzyme activities, respectively. In both methods, DNA is directly extracted from live or permeabilized cells, avoiding the chromatin solubilization and recovery steps that can be challenging. However, these methods require cloning and expressing chimeric fusion proteins that have the enzyme tethered to the protein transcription factor of interest prior to performing the experiment, limiting these techniques from being adapted to in vivo models, patient samples, and histone post-translational modifications.
CUT&Tag vs. CUT&RUN vs. ChIP-Seq
| CUT&Tag | CUT&RUN | ChIP-Seq | |
|---|---|---|---|
| Performed Under Native Conditions? | Yes | Yes | No |
| Chromatin Fragmentation Method | Tn5-based tagmentation | MNase digestion | Sonication |
| Cell Number Requirements | 5,000-500,000 cells | 500,000 cells | 1-10 million cells |
| Sequencing Depth Required * | 2 million reads ** | 8 million reads | 20-50 million reads |
| Integrated Library Preparation? | Yes, uses tagmentation | No, separate library prep required | No, separate library prep required |
| Compatible Targets | Primarily histone modifications, some transcription factors and co-factors | Wide range of histone modifications, transcription factors, and co-factors | Wide range of histone modifications, transcription factors, and co-factors |
| Workflow Length | 1-2 days | 1-2 days | 2-3 days |
* Kaya-Okur et al. Nature Communications (2019) 10:1930
** For less abundant targets of interest, 8 to 10 million reads are recommended
Want to learn more about CUT&Tag? Check out our Epigenetics Podcast with Active Motif scientists Casidee McDonough and Kyle Tanguay to learn the technical details of the CUT&Tag protocol and current developments around this method in our R&D Team.
Before CUT&Tag, There Was CUT&RUN
Inspired by DamID and ChEC, and keeping in mind their limitations, Skene and Henikoff developed the CUT&RUN technique in 2016. CUT&RUN is short for Cleavage Under Targets and Release Using Nuclease. Similar to the ChEC protocol, CUT&RUN uses the endonuclease and exonuclease properties of the MNase enzyme.
For the CUT&RUN protocol, MNase is fused to protein A (pA-MNase) to guide the chromatin cleavage to antibodies bound to the protein targets of interest at their binding sites across the genome. CUT&RUN starts with isolated nuclei from live cells that are immobilized on lectin-coated magnetic beads. The nuclei are then incubated with an antibody specific for the protein of interest and the pA-MNase reagent. The enzymatic reaction is primed by the addition of Ca2+. The protein-DNA complex can be isolated and purified and then used directly for library preparation.
And in the Beginning, There Was TAM-ChIP™
In 2011, Active Motif developed a new method called TAM-ChIP for analyzing protein-DNA interactions. Just like ChIP assays, TAM-ChIP is performed on crosslinked and sonicated chromatin. However, the TAM-ChIP protocol involves using a secondary antibody that is coupled to the Tn5 transposase and NGS adapter sequences. After chromatin capture by agarose beads, Tn5 is activated by Mg2+ to generate the sequencing library index on either side of the protein-binding site, providing higher resolution identification of protein binding sites.
Other similar methods, such as CUT&Tag, are variations on Active Motif’s patented TAM-ChIP technology.
The Rise of CUT&Tag
In 2019, Henikoff’s lab evolved their CUT&RUN method into CUT&Tag.
The CUT&RUN method is good, it can generate high-quality sequencing data from 100-1000 live cells. However, this approach still requires an additional step for adapter ligation prior to library preparation, making it difficult to adapt to single-cell applications.
CUT&Tag uses the Tn5 transposase, just like TAM-ChIP, to simultaneously cleave chromatin and insert the NGS adapters for library preparation. This eliminates a time-consuming step in the workflow and also helps CUT&Tag be compatible with smaller amounts of starting material.
In summary, CUT&Tag analyzes native chromatin like CUT&RUN and uses antibody-guided tagmentation like TAM-ChIP.
What’s Next? Single-Cell CUT&Tag!
In the original CUT&Tag article, the team of researchers from the Henikoff lab adapted CUT&Tag to single-cell analysis. Contrary to CUT&RUN, CUT&Tag is adaptable to single-cell because the whole reaction, from antibody binding to library preparation, occurs within intact cells or nuclei.
For adaption to single-cell analysis, the basic CUT&Tag protocol was slightly adjusted. Instead of using magnetic beads to immobilize the cells, the cells are centrifuged between washing. After the tagmentation step, cells are distributed as single cells in nanowells to be barcoded prior to sequencing.
How Does CUT&Tag Work?

General CUT&Tag Protocol
The CUT&Tag protocol consists of 3 main steps:
- Permeabilization of the native/unfixed cells
- Incubation with the specific antibody
- Targeted tagmentation.
Detailed CUT&Tag Protocol
As mentioned previously, in contrast to TAM-ChIP or classical ChIP assays, cells are not fixed with formaldehyde and are not sonicated in the CUT&Tag protocol. The main steps of the CUT&Tag protocol are outlined below.
- Permeabilization of the native/unfixed cells
- Concanavalin A-coated magnetic beads are washed with the binding buffer.
- Cells are harvested, counted and washed twice with the wash buffer.
- Cells are mixed with the beads to be immobilized and permeabilized with the digitonin-containing antibody buffer.
- Once permeabilized, cells are maintained on ice.
- Incubation with the specific antibody
- Cells are incubated with the primary antibody overnight at 4°C.
- After washing, cells are incubated with the secondary antibody for 1 hour at room temperature.
- In parallel, pA-Tn5-adapter transposomes are prepared. The pA-Tn5 complex is washed with the Dig300 buffer for 1 hour, 3 times.
- NaCl concentration is increased to prevent Tn5 from digesting accessible chromatin.
- Digitonin concentration is lowered to 0.01% to avoid clump formation.
- Targeted tagmentation.
- Cells and pA-Tn5 are mixed in the tagmentation buffer and incubated for 1 hour at 37°C.
- The tagmentation reaction is stopped and DNA is solubilized with the addition of EDTA, SDS, and proteinase K.
- DNA is extracted by phenol/chloroform and precipitated with ethanol.
- DNA is amplified by PCR to insert next-generation sequencing indices.
- To avoid sequencing of PCR duplicates, limit the amplification to 12-14 cycles or fewer.
Following the tagmentation step, the DNA is purified, and the CUT&Tag libraries are ready for sequencing.
CUT&Tag Advantages
CUT&Tag is a newer molecular biology method for investigating protein-DNA interactions and offers some advantages over ChIP-Seq and CUT&RUN.
CUT&Tag is Compatible with Low Cell Numbers
In the publication that first described the CUT&Tag method, the authors went down as low as 60 cells to analyze H3K27me3 profiles across the genome. CUT&Tag is compatible with low cell numbers because pA-Tn5 cleaves DNA directly at the binding site of the antibody and does not require chromatin preparation and sonication steps that can lead to sample loss.
The ability to work with small numbers of cells is an advantage for researchers working on specific cell types, such as rare neuronal populations, pancreatic islets, or stem cells, that are difficult to obtain large numbers of.
CUT&Tag Does Not Require Fixation or Sonication
CUT&Tag is performed on native (unfixed) cells or nuclei, avoiding the need for fixation, chromatin preparation, and sonication steps of standard ChIP workflows. Sonication can be quite challenging to set up and requires specialized equipment that can be expensive. Moreover, over-fixation and over-sonication can destroy protein epitopes preventing them from being immunoprecipitated. Some antibodies work better under native conditions.
CUT&Tag is Fast
Relative to ChIP, which is a multi-step process, CUT&Tag is much faster. Cells are immobilized on magnetic beads and the entire protocol happens in a single tube. The tagmentation step involves chromatin shearing and sequencing library adapter insertion at the same time, which is amenable to high-throughput experiments.
Less Sequencing is Required for CUT&Tag
With CUT&Tag, the chromatin retrieved at the final step of the protocol is in the local vicinity of the binding sites of the protein of interest. The shorter DNA sequences isolated by CUT&Tag means that it doesn’t have the same deep sequencing requirements as ChIP-Seq.
3-5 million reads are sufficient to generate robust data following CUT&Tag protocol, which is approximately 10-fold less sequencing than ChIP-Seq assays, which usually require 30-50 million reads per sample.
To compare the CUT&Tag signal-to-noise ratios with X-ChIP and CUT&RUN, the 3 protocols were run with the same antibody and sequenced to a depth of 8 million reads. The overall results were quite similar between the ChIP-Seq, CUT&RUN, and CUT&Tag methods. However, ChIP-Seq showed higher background and needed deeper sequencing than CUT&Tag.
CUT&Tag is also more easily adaptable to single-cell analyses than ChIP-Seq and CUT&RUN, making it more useful than the other methods for researchers working with heterogeneous populations, including blood, brain, pancreas or liver.
New to CUT&Tag? Let our team of epigenetics experts produce high-quality CUT&Tag results for you in a matter of weeks with our end-to-end CUT&Tag Service. This service will give you the time to focus on the bigger picture and publish faster.
Limitations of CUT&Tag
As promising as CUT&Tag is, it does have some limitations that will likely keep it from completely replacing ChIP-Seq assays.
CUT&Tag’s Native Conditions Are Not Always Suitable
Unfixed/native conditions are not suitable for every experiment. In the original CUT&Tag publication, only histone modifications, NPAT, and CTCF were tested.
Many transcription factors are not abundantly expressed, weakly or transiently bind to DNA, or indirectly bind to chromatin. For these cases, chromatin crosslinking and sonication are mandatory steps to detect protein-DNA interactions.
Furthermore, most ChIP-validated antibodies are validated to work with crosslinking conditions and may not work well with native conditions because the protein epitope availability in crosslinked conditions can be different than in native conditions. Therefore, switching from X-ChIP to CUT&Tag would need thorough antibody validation to be certain of antibody specificity and sensitivity in unfixed conditions.
CUT&Tag is Still New
Although it almost seems like CUT&Tag has been around forever and it is catching on quickly, it was only first published in April 2019. Therefore, there is not yet a very long list of papers containing CUT&Tag results published in peer-reviewed journals.
Although the publication from the Henikoff lab is quite complete, containing a lot of data comparing CUT&Tag to ChIP-Seq and CUT&RUN, the experimental workflow is very different from ChIP-Seq and data from experiments run with this protocol could be difficult to compare to previous ChIP-Seq data, including the results of the ENCODE project.
CUT&Tag Might Introduce Bias
The Tn5 transposase used in CUT&Tag has a high affinity to open-chromatin regions. Therefore, the CUT&Tag approach might be preferentially suited for analyzing histone modifications or transcription factors associated with actively transcribed regions of the genome and not very well suited for the analysis of regions of the genome that are silenced or contain heterochromatin.
Digestion time and pA-Tn5 quantity will need to be carefully optimized for each target to avoid any unspecific tagmentation.
CUT&Tag is a promising technology and time will tell if this method becomes as widely adopted as ChIP-Seq. Like every new protocol, all the flaws have not been detected yet so CUT&Tag optimization must be done with care and every step in the workflow should be checked with suitable controls.
Applications Enabled by CUT&Tag

In the original paper reporting the CUT&Tag technology in the journal Nature Communications, Kaya-Okur et al. applied this new method to various protein targets and histone modifications and various applications. All experiments were performed in the human cell lines K562 and H1.
They first investigated the histone modification H3K27me3, a repressive histone mark, the histone modifications H3K4me1 and 2, which are histone marks associated with active transcription, and RNA polymerase II, which is also a marker of genes being actively transcribed. They compared their results with both the traditional ChIP-Seq workflow and the CUT&RUN protocol, and they found that CUT&Tag generated more sensitive results with less background signal than the other methods.
They also tested CUT&Tag for transcription factors, including the highly expressed factor CTCF and the lower expressed transcription factor NPAT. For NPAT, 1% of the detected binding sites were false positive, meaning that the overwhelming majority of the signal was at real binding sites. CUT&Tag was also able to produce a high-resolution map of CTCF binding sites.
Because the CUT&RUN protocol generated robust data from low cell numbers, the researchers also tested CUT&Tag with low amounts of starting material, including even fewer cells than could be accurately analyzed with CUT&RUN. They analyzed the H3K27me3 modification from a range of 60 cells to 100,000 cells and were able to generate high-quality data from all cell numbers.
Finally, they adapted CUT&Tag to single-cell analysis by slightly modifying the original protocol. Instead of using magnetic beads to immobilize the cells, they centrifuged the samples between the washing steps. After tagmentation, they distributed cells one by one in a nanoplate for barcoding. Single-cell profiles for the H3K27me3 and H3K4me2 modifications were reproducible from one replicate to another. scCUT&Tag was also performed for H3K27me3 in H1 embryonic stem cells and the data allowed the researchers to differentiate the H1 cells from K562 cells.
What’s Next for Antibody-Based Tagmentation Methods?

While CUT&Tag is an exciting new technology in its own right, it is still interesting to speculate about what will come next. Antibody-based tagmentation could be adapted to different methods, including X-ChIP-Seq. The TAM-ChIP method developed by Active Motif uses a secondary antibody directly conjugated to the transposase Tn5 loaded with sequencing adapters. Contrary to CUT&Tag, TAM-ChIP works with formaldehyde-fixed samples, so ChIP-validated antibodies are directly compatible.
Conjugating the Tn5 transposase directly to the primary antibody could increase the sensitivity of antibody-guided tagmentation methods, increase the resolution of detection for the binding site sequence, and reduce the experiment time.
Active Motif has added AbFlex® recombinant antibodies to its antibody collection. These recombinant antibodies contain a transpeptidase Sortase recognition sequence on the Fc fragment of the antibody, which allows the site-specific addition of various labels and tags, including the Tn5 transposase. These AbFlex antibodies could be adapted for use in either the TAM-ChIP or CUT&Tag workflows with Tn5 directly conjugated, allowing for the potential to multiplex the assays and investigate multiple histone modifications or transcription factors simultaneously from the same samples.
Finally, primary antibody-based tagmentation could also be used for MeDIP experiments. This approach would increase both the sensitivity and specificity of this technique by more precisely identifying the methylated sequences with higher resolution.
Summary: New Methods are Breaking Boundaries for Epigenetics Research
Similar to ChIP-Seq and CUT&RUN, CUT&Tag allows the identification of protein-DNA interactions on a genome-wide scale. CUT&Tag is a sensitive method that uses the secondary antibody as an anchor for the Tn5 transposase enzyme fused to protein A to guide cleavage of DNA at the binding site of the target protein and insert next-generation sequencing adapters at the same time.
The low background signal makes this technique suitable for using low numbers of cells as the starting material, at least for histone marks. Moreover, cost-effectiveness and its adaptability to single-cell technology also potentially make this method an option for high-throughput assays.
The antibody-guided tagmentation technology pioneered by Active Motif with the patented TAM-ChIP kit has already led to the development of assays like CUT&RUN and CUT&Tag, and this kind of technology is probably going to be adapted to more and more protocols followed by next-generation sequencing in the future. These new methods are starting to break boundaries and allow researchers to bring epigenetics experiments to a whole new level and will contribute to new and exciting discoveries in the field.
We are looking forward to seeing what comes next. Stay Tuned!
Other Resources You Might Be Interested In:
- Learn about our complete CUT&Tag kits
- End-to-end CUT&Tag service
- Check out our comprehensive ATAC-Seq solutions
- Read our complete guide to chromatin sonication for ChIP
- Listen to our Epigenetics Podcast
- Learn more about the PIXUL® Multi-Sample Sonicator
- Download the ChIP 101 eBook
- Read about how to generate the best ChIP data in every experiment
- Learn how to become a ChIP assay expert
About the author

Anne-Sophie Ay-Berthomieu, Ph.D.
Anne-Sophie was born in the south of France and grew up between the Mediterranean Sea and the Pyrenean Mountains. She grew up as a science fiction fan, leading her to specialize in molecular biology and genetics during graduate school at the University of Lyon, France (secretly hoping her research would give her superpowers!). After living in different places for work, she is back in Lyon, France where she shares her time between her husband, her family, and her friends. During her free time, Anne-Sophie challenges herself with hiking, climbing, racing, and traveling in foreign countries – while waiting for her superpowers to grow!
Contact Anne-Sophie on LinkedIn with any questions, or to tell her about your superpowers.
Related Articles
Next-Gen ChIP-Seq: Genome-Wide Single-Cell Analysis with Antibody-Guided Chromatin Tagmentation Methods
September 5, 2019
New methods are improving the efficiency of epigenomic analysis, enabling analysis of histone modifications at the single-cell level. Many of these methods are variations on Active Motif’s patented TAM-ChIP™ technology, which utilizes antibody-guided tagmentation.
Read More
Complete Guide to Understanding and Using ATAC-Seq
February 9, 2021
The ATAC-Seq method was just published in 2013 and it has already become one of the most common and powerful ways to study chromatin states on a genome-wide level. This article covers what ATAC-Seq is and how to use it in your research.
Read More
<< Back to MOTIFvations Blog Home Page