<< Back to MOTIFvations Blog Home Page
RNA and DNA Methylation Crosstalk – Regulating Transposable Elements AND Cell Fate!

September 27, 2023
Table of Contents:
-
Introduction
-
Does RNA and DNA Methylation Crosstalk Have a Role?
-
The Approach: Proteomic Proximity Labeling to Capture Transposable Element-associated Proteins
-
The Target: The Primate-specific Transposable Element LTR7/HERV-H Locus
-
The Discovery: YTHDC2 and TET1 Crosstalk Regulates Transposable Element Activity and Embryonic Stem Cell Fate
-
Conclusions and Future Research in RNA and DNA Methylation Crosstalk/Transposable Element Regulation
Introduction
The epigenetic silencing of the transposable elements that account for over half of the human genome (Deniz, Frost, and Branco and Padeken, Methot, and Gasser) aids in the vital task of maintaining genome integrity; however, the co-option of some transposable elements as cis-regulatory elements controlling the spatial-temporal expression of genes during eons of human evolution (Hermant and Torres-Padilla) also requires a means of installing activate chromatin states at such sites to allow their activity.
Does RNA and DNA Methylation Crosstalk Have a Role?
While this regulatory mechanism remains incompletely described at the current time, studies had suggested roles for DNA methylation (DNA 5-methylcytosine, 5mC) (Deniz, Frost, and Branco and Padeken, Methot, and Gasser) and RNA methylation (RNA N6-methyladenosine, m6A) (e.g., Liu et al.) in silencing transposable element activity. Interestingly, research in human cancer cell lines has provided proof that crosstalk between m6A and 5mC could regulate chromatin accessibility and transcription of enhancers and promoters (Deng et al.); therefore, researchers led by Yarui Diao (Duke University) explored the relevance of RNA and DNA methylation crosstalk in regulating transposable element activity in human embryonic stem cells (Sun et al.).
The Approach: Proteomic Proximity Labeling to Capture Transposable Element-associated Proteins
The authors of this fascinating study developed a targeted proteomic proximity labeling approach to capture transposable element-associated proteins in human embryonic stem cells to reveal the epigenetic mechanisms controlling the repression/activation of transposable elements. They named this CRISPR-based transposable element-centric proteomics approach “CARGO-BioID” (“chimeric array of guide RNA oligos” with proximity-dependent “biotin identification”), which helps to overcome the array of technical challenges that had faced researchers hoping to identify the effectors controlling the activity of endogenous transposable element sequences.
The Target: The Primate-specific Transposable Element LTR7/HERV-H Locus
The team chose to study the primate-specific transposable element LTR7/HERV-H locus in human embryonic stem cells, given the presence of active histone marks (such as histone H3K4me1, H3K4me3, and H3K27ac), the function of these loci as enhancers, promoters, pluripotency factor binding sites, and topologically associating domains boundaries (Leung et al., Zhang et al., Glinsky, 2015, and Gorkin, Leung, and Ren), and the loss of active histone marks at these loci during differentiation (Xie et al. and Leung et al.).
After the application of the CARGO-BioID protocol in human embryonic stem cells, mass spectrometry analysis then identified nearly 150 candidate proteins that displayed elevated levels of enrichment at LTR7/HERV-H loci; correspondingly, putative LTR7/HERV-H loci-associated proteins participated in functions such as histone modification, chromosome organization, transcription initiation, messenger RNA processing, and pluripotency factor signaling.
The Discovery: YTHDC2 and TET1 Crosstalk Regulates Transposable Element Activity and Embryonic Stem Cell Fate
Excitingly, Sun et al. discovered that the YTHDC2 (YTH N6-Methyladenosine RNA Binding Protein C2) protein, which functions as a “reader” of m6A, became bound to LTR7/HERV-H loci through interactions with m6A-modified HERV-H RNAs to limit their transcriptional and epigenetic silencing. Previous research had demonstrated that binding of the related YTHDC1 protein to RNAs associated with transposable element loci induced their degradation or recruited histone modifiers to promote epigenetic silencing (Jambhekar, Dhall, and Shi).
Unexpectedly, they discovered that YTHDC2 recruited the 5mC-demethylase TET1 (Tet Methylcytosine Dioxygenase 1) to remove the 5mC modification from LTR7/HERV-H loci and prevent epigenetic silencing via active DNA demethylation. Finally, the team additionally revealed a role for this interaction in the regulation of cell fate by discovering that YTHDC2 normally acted to inhibit the ability of human embryonic stem cells to differentiate toward neuronal lineages by acting on LTR7/HERV-H loci.
Conclusions and Future Research in RNA and DNA Methylation Crosstalk/Transposable Element Regulation
Overall, the findings presented here have expanded our knowledge regarding the regulatory crosstalk existing between the epitranscriptome (RNA m6A) and the epigenome (DNA 5mC) by providing evidence for the critical nature of YTHDC2 and TET1 crosstalk in regulating transposable element activity and human embryonic stem cell fate. Furthermore, their development of the CARGO-BioID technique may open multiple research lines to further our understanding of transposable element function and regulation in a wide range of biological processes and human diseases.
For more on how RNA and DNA methylation crosstalk in human embryonic stem cells controls transposable elements and cell fate, see Nature Genetics, July 2023.
About the author

Stuart P. Atkinson, Ph.D.
Stuart was born and grew up in the idyllic town of Lanark (Scotland). He later studied biochemistry at the University of Strathclyde in Glasgow (Scotland) before gaining his Ph.D. in medical oncology; his thesis described the epigenetic regulation of the telomerase gene promoters in cancer cells. Following Post-doctoral stays in Newcastle (England) and Valencia (Spain) where his varied research aims included the exploration of epigenetics in embryonic and induced pluripotent stem cells, Stuart moved into project management and scientific writing/editing where his current interests include polymer chemistry, cancer research, regenerative medicine, and epigenetics. While not glued to his laptop, Stuart enjoys exploring the Spanish mountains and coastlines (and everywhere in between) and the food and drink that it provides!
Contact Stuart on Twitter with any questions
Related Articles
LINE-1 Transposable Elements: Can they Provide Liquid Biopsy Biomarkers for Early Cancer Detection and Management?
April 4, 2023
Detection of LINE-1 ORF1p in liquid biopsies has been proposed as an effective means of diagnosing cancer early and improving therapeutic outcomes. In this article, we take a look at two related, innovative studies describing how LINE-1 transposable elements may provide new biopsy biomarkers whose detection will improve the early diagnosis and long-term management of many cancers.
Read More
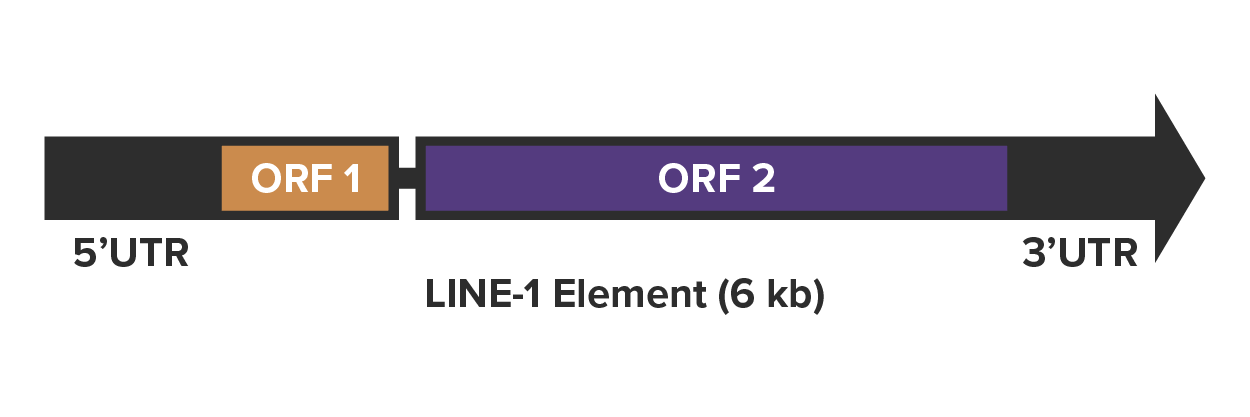
LINE-1 Elements: Walk the LINE-1
September 13, 2022
While LINE-1 elements have been vital for fueling genetic variation and human evolution, their repression and that of other transposable elements is crucial to maintain homeostasis in cells so that our genomes continue to walk the line! Read how rogue LINE-1 expression is a target for combating its role in disease and aging.
Read More
<< Back to MOTIFvations Blog Home Page