What Are HDACs?
Histone deacetylases (HDACs) are a class of enzymes responsible for removing acetyl groups from lysine residues contained in proteins. These acetyl groups have been added by another class of enzymes called histone acetylases (HATs). HDACs and HATs function together to maintain the acetylation status of proteins in the cell, which can contribute to protein stability, protein-protein interactions, gene regulation, and more. For more detailed information on HDAC classifications and function, please see this review.
Histone Acetylation and Chromatin Structure
You could be forgiven for assuming that HDACs function to remove acetyl groups only from histone proteins, but you'd be incorrect. It is well documented now that HDACs remove acetyl groups from histone and non-histone proteins alike. But if we focus only on histone proteins for a moment, we can generally say that HDACs cause a condensed chromatin structure and transcriptional repression. Acetylation of the lysine residues located on histone tails results in a relaxed chromatin structure and transcriptional activation. HDACs remove these acetyl groups, and push chromatin structure to a 'closed' status.
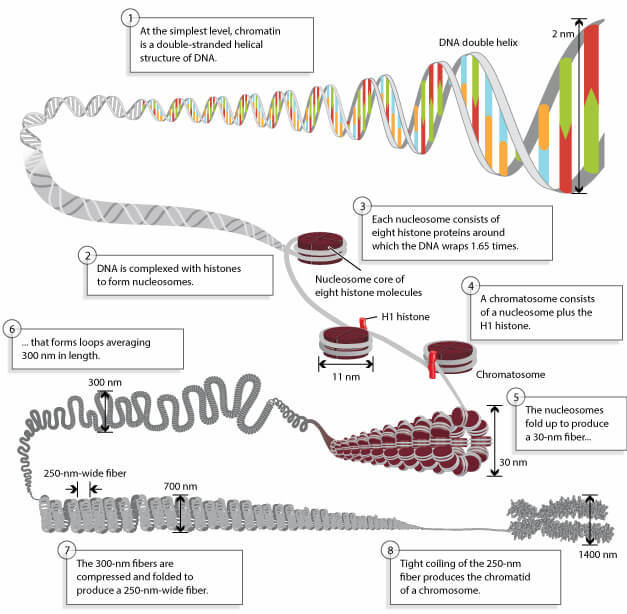
Chromatin consists of DNA wound around histone proteins. Acetylation of histones affects how 'relaxed' and accessible the DNA is. Adapted from Pierce, Benjamin. Genetics: A Conceptual Approach, 2nd ed. All rights reserved.
HDACs in Cancer
Aberrent HDAC expression has been identified across a wide variety of cancers, as has been published in dozens of articles. In general, increased HDAC expression can be used as a biomarker for cancerous tissue, a negative prognostic indicator, and a predictor of decreased survival. However, there are also reports of increased HDAC expression correlating with improved prognosis, and evidence that some HDACs serve as tumor suppressors. HDAC inhibitors (HDACis) have sucessfully contributed to treatment of various cancers and non-cancerous diseases (e.g. rheumatoid arthritis), but based on the conflicting functions for HDACs in cancers, one might not be surprised if chronic HDACi treatment someday resulted in therapy-induced tumorigenesis.
Tools for HDAC Analysis
HDAC Activity Assay
A simple first step to analyzing HDACs is to examine HDAC activity in cellular or nuclear extracts from your cells. This allows you to see global changes in enzymatic activity.
ELISA Assays
You may also want to examine the substrates, proteins being deacetylated. ELISA assays and Western blots using validated HDAC antibodies and HDAC proteins can provide a quick, global view of acetylation at a specific site on a specific protein.
Mod Spec Service
For a more high throughput analysis of which proteins are being deacetylated, perhaps in response to a new HDACi that you are screening, you may want to consider the Mod Spec® Service.
ChIP-seq
To go one step further, you can identify the genomic locations where a specifically acetylated protein is binding by performing ChIP-seq with a ChIP-validated antibody.
RIME Analysis
And last, once you've identified the speific acetylation sites and genomic binding sites, RIME analysis can identify HDAC-interacting proteins that may be driving HDAC action in a site-specific manner. No matter what tool you use, HDAC analysis opens up a world of new hypotheses regarding chromatin structure, protein-protein interactions, and gene expression regulation to elucidate the mechanisms of your disease model.

