<< Back to Resources Home Page

Download Our Scientific Posters
Active Motif’s team of expert scientists travel around the world presenting data at conferences and scientific meetings. Here’s a list of some of our posters we presented for you to download if you weren’t able to attend.
Generation of Targeted Antibody Conjugates Using Sortase and the AbFlex® Technology Platform
This presentation from the American Association of Immunologists (AAI) 2024 highlights the development of a novel antibody conjugation technology, the AbFlex platform, a versatile epitope tagging system for the carboxyl terminus of recombinant mouse and rabbit immunoglobulin heavy chains. It consists of a tandem array of three tags that enable generation of a wide array of user-defined antibody conjugates.
Download the posterA Universal Spike-In Normalization Strategy for CUT&RUN, CUT&Tag, and ATAC-Seq
This poster was presented at AACR 2024, describing the development of unique reagents for assay normalization. The use of Spike-In controls as a means of normalization in NGS based assays is shown to be not only useful, but in some cases necessary to detect true differences in gene expression or protein occupancy. Data is presented comparing Spike-In normalization with techniques using Drosophila nuclei.
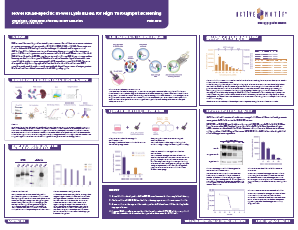
Download the posterA New HTS ELISA for Monitoring NRAS Activity to Expedite Drug Discovery
Oncogenic mutations in the KRAS, HRAS, and NRAS isoforms of the RAS gene family alter GTP binding and hydrolysis leading to errant over activation of the MAPK/ERK pathway. While the total of all such RAS mutations drives some 20-25% of all human cancers, each isoform tends to favor particular mutations that are cancer specific. NRAS mutations are seen in about one-fifth of cutaneous melanomas and patients with NRAS mutation melanomas have a poorer prognosis due to the high aggressiveness of RAS mutant tumors, a lack of efficient targeted therapies, or rapidly emerging resistance to existing treatments. In this AACR 2024 poster, we describe the development of a highly specific and sensitive ELISA for determining the active NRAS protein in high throughput compatible (HTS) format.
Download the posterUtilization of MBD-Seq to Elucidate Differentially Methylated Regions in cfDNA Across Healthy and Cancer Patients
The 5-Methylcytosine (5-mC) epigenetic DNA modification alters gene expression and genome metabolism, and its deregulation is associated with many human diseases. Cell-Free DNA (cfDNA) released from apoptotic cells has recently gained attention, as mutations or epimutations (e.g. 5-mC variations) detected in cfDNA were shown to have high diagnostic potential to assess the presence, stage and outcome of several cancers. However, 5-mC detection in cfDNA remains a challenge as input material is often very scarce. MBD-Seq is a method that leverages the ability of Methyl-CpG-binding domain protein 2 (MBD2) to capture and detect highly methylated regions of DNA genome-wide. This AACR 2024 poster demonstrates use of MBD-Seq for the detection of aberrant methylation patterns in cfDNA from healthy and diseased patients, finding it to be specific and sensitive, revealing striking differences in methylation of key tumor-associated genes.
Download the posterHigh-throughput Genomic Analysis of Urine Samples Using Pixelated-Ultrasound to Improve Urine Biopsy Workflow
This AACR 2024 poster describes an improvement to sample preparation of liquid biopsy from cancer patients for sequencing. The advantages of a novel, pixelated ultrasound technology significantly improves sample preparation in a fast, economical, high-throughput manner for both low and high cell numbers. This method has the potential for integration in automation workflows and across large cohorts.
Download the posterNovel KRAS-specific In-well Lysis ELISA for High Throughput Screening
This poster was presented at AACR 2023, describing the development of an activation state specific assay for KRAS, as well as Active Motif’s novel Single B-cell cloning technology for development of the recombinant antibody which provides the key specificity for the assay.
Download the posterNovel Epigenetics Technology for High-Throughput Processing of Limited Samples to Study Cancer Using Cavitation-based Pixelated Ultrasound and Tagmentation-indexing ChIP-Seq
This poster was presented at AACR 2023 and describes two novel technologies. One workflow applies pixelated ultrasound to enable sample preparation critical to a range of downstream applications. The second method presented is our new Tagmented, Indexed and Pooled ChIP-Seq (TIP-ChIP), a novel epigenetics assay to achieve high-throughput, multi-mark ChIP-Seq.
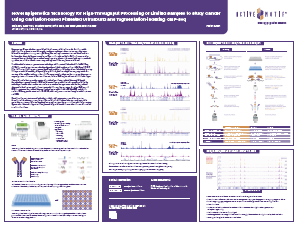
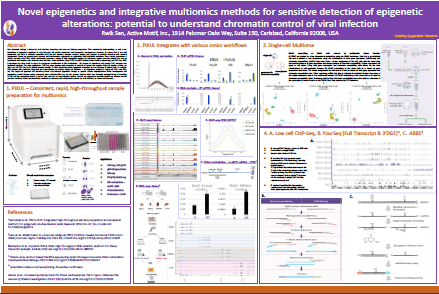
Download the posterNovel Epigenetics and Integrative Multiomics Methods for Sensitive Detection of Epigenetic Alterations: Potential to Understand Chromatin Control of Viral Infection
This poster was presented at the NIH Chromatin Control of Viral Infection Conference, 2022. The presentation features the incorporation of the PIXUL® Multi-Sample Sonicator as an intregal part of sample preparation in single-cell multiomic analysis via ATAC-Seq and RNA-Seq.
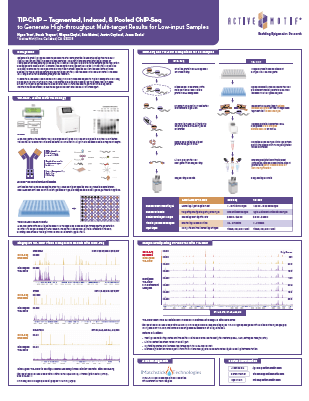
Download the posterTIP-ChIP: Tagmented, Indexed and Pooled ChIP-Seq to Generate High-Throughput, Multi-Target Results for Low Input Samples
Winner of a Reviewer’s Choice award, this poster was presented at the American Society for Human Genetics (ASGH), 2022. TIP-ChIP (Tagmented, Indexed, and Pooled ChIP-Seq) is a new method developed to rapidly complete ChIP-Seq on 96 samples for multiple epigenetic marks.
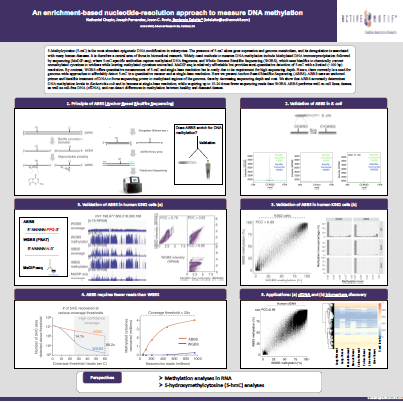
Download the posterAnchor-Based Bisulfite Sequencing (ABBS), Enrichment-Based Nucleotide-Resolution Approach to Measure DNA Methylation
Recently presented at Cold Spring Harbor, 2022, this poster presents Anchor-Based Bisulfite Sequencing (ABBS), developed as a method to determine DNA Methylation levels at single-base resolution, requiring up to 20X fewer reads than WGBS, with reduced sequencing depth and cost, and suitable for cell lines, tissue, and cfDNA sample types.
Download the posterNew High-Throughput Methods for the Epigenomic Characterization of Human Disease States
The new methods described in this poster, such as the PIXUL 96-well sonicator, transposase-assisted multiplex ChIP (TAM-ChIP), high-throughput ChIP, low cell number ChIP, and ChIP from FFPE tissues, are making it easier to characterize the epigenomic profiles associated with human diseases.
Download the posterEpi-Seq: Epigenome Landscape Mapping for Clinical and Drug Discovery Settings
This poster describes Epi-Seq, a complete experimental solution that will dramatically reduce the technical expertise and touch-time needed to perform ChIP-Seq, while also increasing consistency and throughput.
Download the posterNovel High-Throughput Tools for Detecting Age-Associated Epigenetic Changes in Liquid Biopsies
This poster demonstrates the applications of Active Motif’s high-throughput approaches to detect epigenetic alterations in human serum samples from healthy young and old individuals.
Download the poster