<< Back to Resources Home Page
Enabling Epigenetic Analysis of Liquid Biopsy Samples

Liquid biopsies are changing the ways physicians and researchers diagnose and treat many different diseases. Improvements to research tools and analysis methods are making it possible to perform genetic and epigenetic tests on the small amount of material present in most common liquid biopsy samples. These newer tests can give more information to doctors faster than previous approaches, allowing more rapid diagnostics and better treatment strategies. Liquid biopsies are less expensive and less invasive than solid tissue biopsies, so they are also generally better for the patient.
However, analysis of liquid biopsies, and epigenetic analysis in particular, is still relatively new and this is a very active research area. Clinical researchers are using liquid biopsy samples to identify and validate novel biomarkers, such as DNA methylation profiles, to improve how quickly they can detect diseases and also how they can treat their patients.
What is a Liquid Biopsy?
A liquid biopsy is a non-invasive method to obtain biological samples from a patient. Liquid biopsies are most commonly blood, plasma, or serum samples, but they can also be derived from other biological liquids.
Liquid biopsies can be analyzed for many different types of biomarkers and they are generally used in diagnostic or prognostic tests to monitor disease progress, responses to treatments, or to look for early warning signs of diseases.
Because liquid biopsies are non-invasive, usually just requiring a blood draw, multiple samples can be taken frequently over a short period of time. This approach is preferred over solid tissue biopsies in many cases because it allows generation of multiple data points, providing more complete information to physicians regarding a patient’s health and therefore allowing doctors to more closely monitor treatments and make any necessary changes as quickly as possible.
The most common applications of liquid biopsies are currently diagnostic tests for human cancers. In these cases, the liquid biopsy samples are analyzed for the presence of circulating tumor cells (CTCs) or circulating cell-free DNA (cfDNA) containing mutations associated with cancer. Assays are also being developed to identify and detect epigenetic profiles associated with cancer and other human disease states in liquid biopsies.
Investigating Epigenetics of Liquid Biopsy Samples
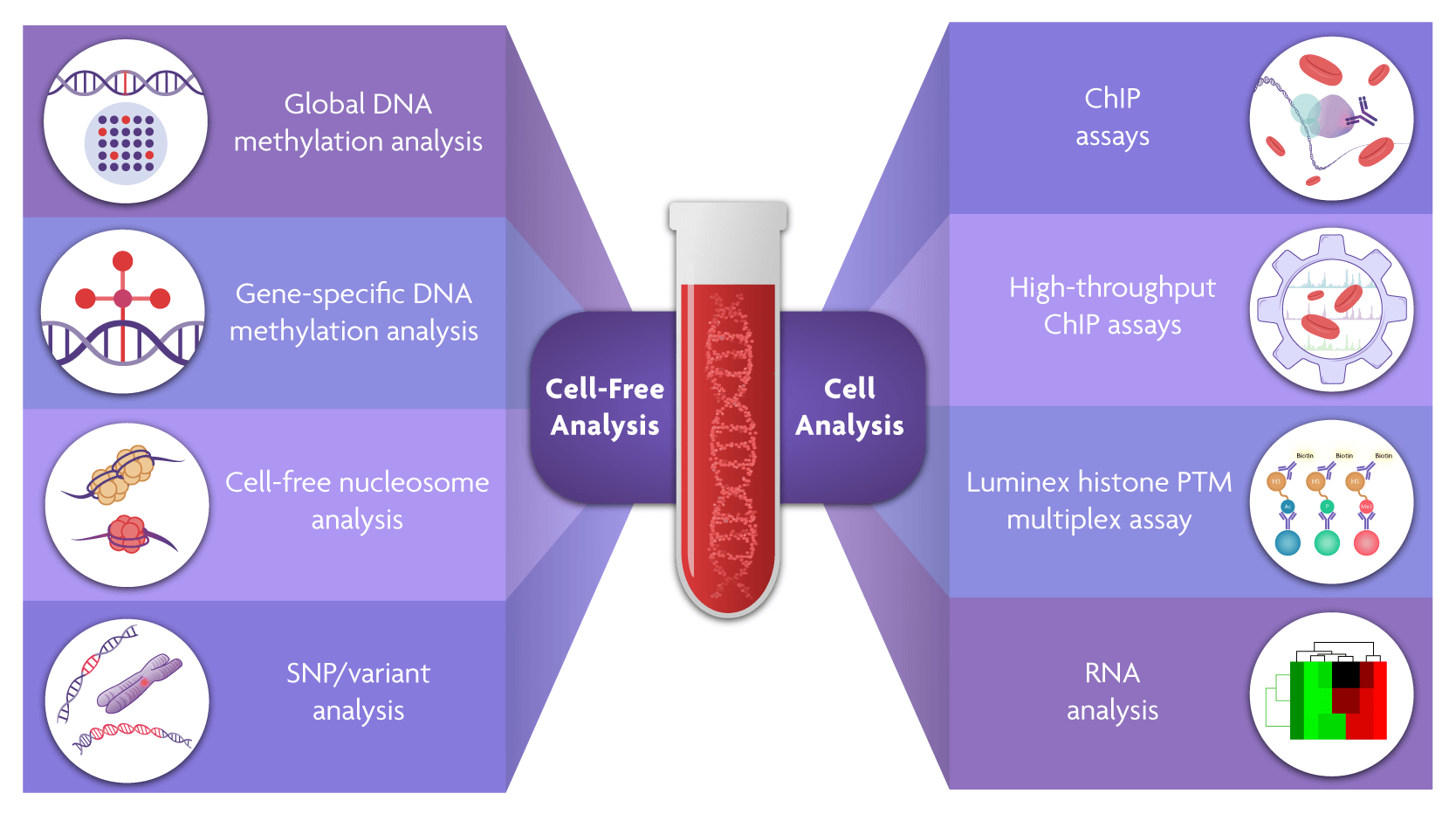
Epigenetic analysis of liquid biopsies has so far primarily focused on investigating the DNA methylation status of cell-free DNA and the presence and modifications of circulating cell-free nucleosomes.
Methods to Analyze DNA Methylation in Liquid Biopsy Samples
DNA methylation assays with liquid biopsies generally start will a sample preparation protocol to isolate the cell-free DNA in plasma or serum samples. The Cell-Free DNA Purification Kit makes this part of the experiment easy and efficient. cfDNA that has been shed into the blood is thought to be enriched for regions of the genome that contain disease biomarkers. Therefore, it is important to specifically isolate cfDNA, rather than total genomic DNA from leukocytes, to ensure the most sensitive and specific detection of biomarkers in cell-free DNA.
After the cfDNA has been purified, the next type of assay performed depends on the experimental goals. If values for the global DNA methylation levels are required, a high-throughput ELISA assay is usually performed. This type of global assay is appropriate for high-throughput analysis of large or widespread changes in DNA methylation, but may not be sensitive enough to detect methylation changes at a few specific regions in the genome.
If the goal is to determine the methylation status at specific regions of the genome, assays like methylated DNA immunoprecipitation (MeDIP) or methyl binding domain enrichment assays are more appropriate. To look at methylation on a genome-wide scale, then methods like MeDIP-Seq or reduced representation bisulfite sequencing (RRBS) would be required. These methods allow researchers to investigate changes in DNA methylation levels on a genome-wide scale without the costs of sequencing the entire genome.
Many researchers use a combination of several different DNA methylation assays for the most complete analysis of their liquid biopsy samples. For example, clinical researchers often start with a high-throughput global DNA methylation assay on their entire cohort of hundreds of samples. They then use the results of that ELISA-based assay to determine which samples to investigate further with MeDIP kits, or RRBS services.
Methods to Detect and Measure Circulating Cell-Free Nucleosomes
Another approach to analyzing the epigenetic profile of liquid biopsy samples is to measure the levels of circulating cell-free nucleosomes, which are a complex containing DNA wound around histone proteins. Increased levels of circulating nucleosomes in serum are associated with several different human disease phenotypes. Therefore, cell-free nucleosomes have become increasingly established as a novel class of epigenetic biomarkers.
Performing ChIP Assays on Blood Samples
Traditional chromatin immunoprecipitation (ChIP) assays can also be performed on peripheral blood mononuclear cells (PBMCs) derived from whole blood samples. The cell types present in PBMC samples are generally more difficult to work with than other sample types because they are smaller and harder to lyse.
The ChIP-IT® PBMC kit solves this problem by including a lysis and chromatin preparation protocol that was specifically developed for these cell types, making ChIP assays with PBMC samples as easy and efficient as possible.
Resources to Learn More About Epigenetic Analysis of Liquid Biopsies
Epigenetics is complicated, and sometimes it feels difficult to get started or keep up with this fast-moving field. We’ve got you covered! Our growing list of educational resources makes it easier than ever to become an epigenetics expert.
Featured Liquid Biopsy Resource
Epigenetics changes, such as DNA methylation and levels of cell-free nucleosomes, are known to be associated with aging. Liquid biopsies, using samples such as plasma or saliva, have become an increasingly common approach to investigate epigenetic changes in human patients.
This scientific poster demonstrates the applications of Active Motif’s high-throughput approaches to detect epigenetic alterations in human liquid biopsy samples from healthy young and old individuals.




